Path-o-gen is a handy tool from Andy Rambaut’s group ‘for investigating the temporal signal and ‘clocklikeness’ of molecular phylogenies’. It is a common pre-cursor to the more involved BEAST analysis, to see whether there is any clock-like signal (i.e. SNPs accumulating with time) in the phylogeny.
I wanted to do this today to look at temporal signal within an outbreak phylogeny, and I ran into a few problems. This is one of those ‘write it down before I forget’ posts.
So, I have a tree with a bunch of samples on. I have a bunch of dates associated with those samples (day the person’s sample was taken). You can manually enter the sample dates, or Path-o-gen has various ways to ‘guess the dates’. It does this based on the taxon label (i.e. your sample name from your tree).
I tried a bunch of ways of feeding the information into path-o-gen via the ‘guess dates’ function, and this is the one that works. There is almost certainly a better way of doing this, if you know it, please leave a comment!
Essentially, you need to rename your taxon labels with the ‘year proportion’. This is, for example, 2015.704109589041096 for 15th September 2015 (that date is 0.70 of the way through the year). Here is a script to take in a bunch of sample ids, and output the sample id and the year.proportion.
Then, you use the awesome newick utils to rename the taxon labels on your newick format tree. This is a command line tool that takes your tree and the tab separated output that the above script will produce and spits out a tree with the taxon labels replaced with the year proportion.
Then, you can just load this into path-o-gen, press ‘guess dates’, leave everything as default and it will have successfully populated your date/height columns. Check the height columns are all sensible, and the maximum height is not more than your oldest sample. Then, the fun part!
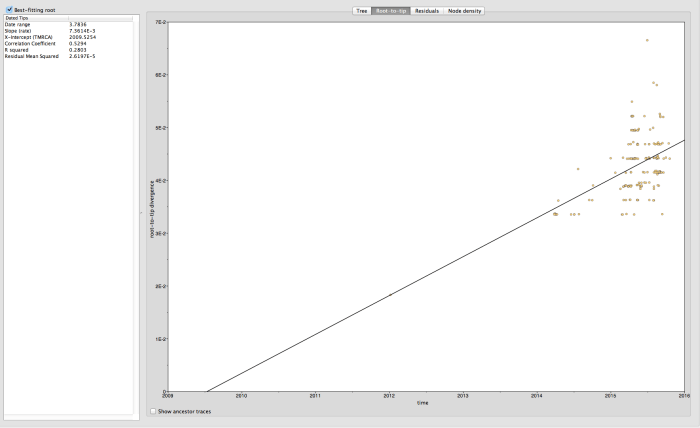
Not bad – correlation co-efficient of 0.53
One thing with this approach (naming the tip labels after the date) is that, if you have two samples taken on the same day (not uncommon in an outbreak scenario, but maybe less of a problem for broader gen epi studies), the taxon labels will be the same and path-o-gen will baulk. Therefore, the script above checks if you have two date proportions the same, and if you do, adds a random number that should be less than 0.0000099, which in terms of proportion of a year is around 30 seconds. This means your numbers won’t be identical, but your result should not be affected. Bit hacky, but it works (i think). If you didn’t read this far down, and this somehow screws your results, then maybe you shouldn’t just run code you find on blog posts without looking at it 🙂






