This is a post by Lauren Cowley.
I have just spent one amazing month in Australia with Scott Beatson’s group at the University of Queensland. I was lucky enough to be awarded the Presidents fund grant by the SGM that enabled me to make the visit and am pleased with all the productive collaborations and work I’ve achieved. While I was there I was looking further into my plans to perform TraDIS as well as learning about and using some of the Beatson group’s genome comparison tools.
They have developed lots of really ace tools there but I was working with BRIG (http://www.biomedcentral.com/1471-2164/12/402, http://sourceforge.net/projects/brig/), EasyFig (http://bioinformatics.oxfordjournals.org/content/early/2011/01/28/bioinformatics.btr039, http://sourceforge.net/projects/easyfig/) and SeqFindR (https://github.com/mscook/SeqFindR) especially.
Firstly I used BRIG, the Blast Ring Image Generator, which is a really nice and easy to use whole genome comparison tool that gives a cool circular image that is easily interpreted. Similar to many other tools, it uses BLAST and CGView 0to generate visual comparisons and colours regions of high sequence identity. What differentiates BRIG from tools like circos that may produce similar plots is that it can be executed very easily with a graphical user interface and therefore is very appealing to those not so familiar with the command line. The BLAST is done against an inputted reference that can either be a singular genome or, as in my case, a multifasta of multiple genomes to show similarities within a group that do not have a unifying reference genome.
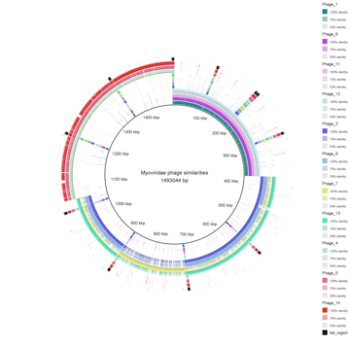
The above BRIG image was generated to show the similarities within a group of T4-like bacteriophages that infect Escherichia coli O157, the central multifasta reference included all the phages in the group in the order they are listed on the right hand side of the image. This order is also the order of genomes in the outer concentric rings. Each region of the multifasta shows the hits to that genome with other genomes in the outer rings, including a hit back from itself. This image shows us that the group of phages can be further sub-divided into 3 groups of phages based on their sequence similarity. The first group consisting of phages 1, 8, 11 and 12, the second consisting of 3, 6, 7 and 13 and then the third consisting of 4, 5 and 14. The final ring has been allocated to mark the tail coding region of the phages that is shared with all members of the Myoviridae phage group. The BRIG image also shows us that there is very little sequence similarity between the groups.
I wanted to do further analysis of the accessory regions within each sub-group of phages and luckily for me the Beatson group had two tools that when used in collaboration would make some really nice figures for this also. These tools were EasyFig and SeqFindR, both python applications. EasyFig like BRIG also comes with a nice graphical user interface and produces easy to view Artemis based figures for regions of a genome that you want to compare linearly. These figures can also be interactively coloured. SeqFindR is a command-line tool that produces plots of hits in an assembled genome, and/or consensus mapping, for a multifasta file of genes of interest that you put together.
The above figure is a combination of an EasyFig image, blue and red arrows with blocks of similarities between, overlaid on a SeqFindR plot, accessory gene hits represented in columns. The EasyFig image represents the whole genomes of phage 4, 14 and 5 with the accessory genes coloured in red and regions of high genomic similarity represented by linking blocks. The SeqFindR image has then been dissected and aligned to the corresponding accessory regions to show the presence or absence of that gene in each phage for group 3 of the Myoviridae phage. This visualisation gives a clear and easily interpretable image for looking at accessory genes that can characterise their presence or absence within a group of genomes and show an annotation for that gene.
The use of these tools for genome comparison has been very useful for my work and I’m sure could be widely used by many others who are trying to do similar things. I had a fantastic time in Oz with the Beatson group and they made sure I had brilliant weekends as well as working hard during the week. I tried my hand at surfing and was pretty pleased that I successfully managed to stand up but can’t say I got much further than that. Thanks guys for an awesome trip to the land down-under!!
From left: Bryan Wee, Majed Alghoribi, Nabil Alikhan, Me and Scott Beatson









Where is a photo of all of us? Where is the GoPro footage of you surfing? Where is the complaint about SeqFindRs complicated install. Anyway, glad to see that you found the tools useful.
from Lauren
Hah, there is no photo of all of us :-L and no GoPro footage as yet. Haha I also left off the complicated install four letter words (I jest)! Very useful and clever tools indeed!!