Once your phylogenetic tree gets above 50-100 strains it can become a bit unwieldy to analyse. One nice way of getting around this is to annotate your tree with relevant info e.g. serotype, phage type, source, date, location etc.
A while ago, Tim wrote a script to annotate NEXUS format trees for viewing in e.g. FigTree. I recently adapted the script to add in colour as well.
There are a few vignettes within this post that might be of interest to different people.
- How to assign different names to taxa in NEXUS trees
- How to colour taxa in NEXUS trees
- Some example colour schemes for use in figtree NEXUS trees
1. and 2. are both covered in this script. As usual, any questions, just ask. This script has a few things that are specific to the layout and structure of our own data, so you will need to be able to fiddle around in python to adapt it for your own data.
In terms of the actual colours, I looked for an explanation of the colouring system that figtree uses but with no luck, so I reverse engineered a colour scheme. To save you the minor hassle of doing that, here is a colour scheme and the numbers you need to recreate it in figtree.
These colours are [‘16737895’, ‘10040065’, ‘10066177’, ‘16777012’, ‘6750055’, ‘39220’, ‘39322’, ‘3407872’, ‘6711040’, ‘205’, ‘6684775’, ‘16738048’]
I also made a red to green one, although the resolution is a bit limited (i.e. the difference between some of the colours is minimal). The numbers for that one are in the script.
Then, when I run the script on the tree – all the info we need to analyse it is written on (too small to read, PPI issues) and the taxa are coloured according to month of isolation, allowing us to quickly spot patterns.
If anyone has any insight into the colour scheme used by Figtree, I would love to hear about it.
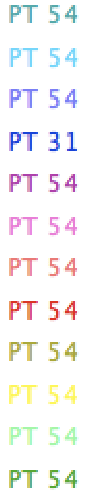








I convert to my tree figures to SVG and then annotate, recolour using svg directives.
Following your script to modify the nexus file to add the color tag for each taxa independently, I saw that figtree is also able to identify colors in hex code, making it easier to select which colors use in the tree, but still the modification can only be done directly over the nexus file following some criteria (the month where the sample was taken in your example). Because I still have no clue how to use the color scheme option in figtree.