TL;DR
- I have run Torsten’s MLST script on the NCTC PacBio genomes, here are the results.
I’m a massive fan of the NCTC3000 project, a collaboration between the National Collection of Type Cultures and the Wellcome Trust, to PacBio sequence 3000 bacterial type strains from the NCTC collection.
One of the reasons I like the project is that it seems quite open. All the data is being made available on a rolling basis here, and there are links to batch download all the assemblies rather than just the raw data. Don’t get me wrong, having the raw data is great, but PacBio assembly is quite computationally intensive (for now), so having the assemblies is fantastic!
Anyway, I have wanted to get the MLST STs of the Salmonella genomes that NCTC3000 have sequenced for a while. When I heard about Torstens neat MLST package for getting the ST from an assembly in a single command, I thought it would be neat to get STs for ALL the thingz.
So, as an approximate protocol,
- I downloaded all the genomes from here
- Since there is a mix of gffs and EMBL files in there, I needed to install python GFF parsing from BCBio. I already had EMBL parsing from BioPython SeqIO
- I ran this script to pull out the contigs in the embl and gff files as fastas
- Finally, I ran Torsten’s MLST package on all the fasta files, without specifying a scheme. It tries to guess which scheme to use by blasting the genome against all the alleles and choosing the scheme with the most hits.
The results are quite interesting, available here for your viewing pleasure. A few caveats – there is currently no minimum threshold for the ‘guess the scheme’ algorithm. Therefore, there will be some spurious comparisons, e.g. something NCTC call Actinobacillus lignieresii was compared against the ecloacae MLST scheme. However, no ST was found, so no harm done. Secondly, these results have only been the most minimal of sanity checking. If you are interested in any of these genomes, download it yourself, convert to fasta and run the mlst.
Some cursory analysis – ecoli is by far the most sequenced genome. I’m looking forward to seeing what people find out about ecoli with 230 reference quality genomes! 60 of those genomes are ST10, which, from a quick google seems to include some E. coli O78:H10 enteroaggregative E coli, which are also associated with urinary tract infections.
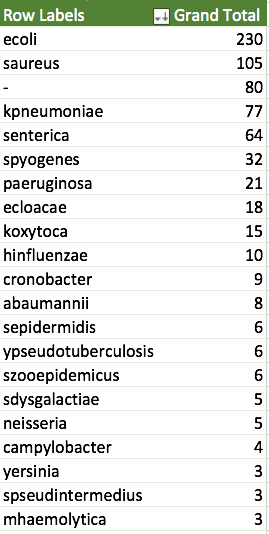
Also, if we assume that STs returned as ‘-‘ are novel STs, then it is interesting that 80% of the Klebsiella oxytoca are novel STs, half of Enterobacter cloacae are also novel STs.






