Adapted by Lauren Cowley from original PHE training material by Philip Ashton and Tim Dallman
Answers at the end
Salmonella Enteritidis PT14b outbreak exercise
Bioinformatics training, interpretation of phylogenies
Prepared by the Gastrointestinal, Emerging and Zoonotic Infections department and the Gastrointestinal Bacteria Reference Unit, Public Health England, Colindale, London.
Aim of Session
To develop an understanding of how whole genome sequencing can help in the investigation of outbreaks of gastrointestinal infection.
Learning objectives
By the end of this exercise participants should:
- Understand the extra value of WGS based typing over traditional typing techniques and the epidemiological implications of this.
- Be able to interpret phylogenetic trees in the context of GI outbreaks.
- Have reflected on the impact of WGS on their current practice.
Summary
This scenario is based on a national outbreak of Salmonella Enteritidis PT14b (Rapid draft sequencing and real-time nanopore sequencing in a hospital outbreak of Salmonella. J Quick, P Ashton et al. Genome biology 16 (1), 114). However, this is the Hollywood remake of the British original and the details vary from reality.
This scenario provides:
- Experience interpreting whole genome sequencing data in the context of an outbreak
- An opportunity to put existing typing techniques into the context of WGS
- An example of the importance of ‘time, place and person’ context in the interpretation of WGS
It also considers practical issues in the management of an outbreak including communication, data management, roles and responsibilities and the complexities of national outbreak investigations.
Inject 1a: The first cases
It is Monday 9th June 2014. The previous week the national reference lab received a phone call from a hospital reporting an unusual number of Salmonella Enteritidis

Sure enough, come Monday morning, the West Midlands cases trigger an exceedance (see Fig 1, Table 1)
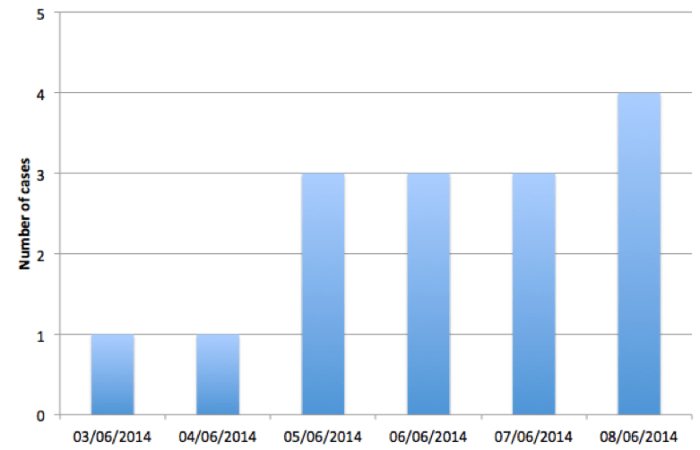
Figure 1: Epidemic curve of cases reported in the exceedance of 09/06/2014
Table 1: Line listing of patients making up the exceedance of PT14b. Hospital or community refers to whether or not the patient’s onset date is consistent with acquisition during hospital stay.
| Region | Sample date | Age | Hospital or community | Travel |
| West Midlands | 03/06/2014 | 2 | Community | Spain |
| West Midlands | 03/06/2014 | 62 | Hospital | No |
| West Midlands | 04/06/2014 | 41 | Hospital | No |
| West Midlands | 05/06/2014 | 23 | Hospital | No |
| West Midlands | 05/06/2014 | 58 | Hospital | No |
| West Midlands | 05/06/2014 | 67 | Hospital | No |
| West Midlands | 06/06/2014 | 45 | Hospital | No |
| West Midlands | 06/06/2014 | 62 | Hospital | No |
| West Midlands | 06/06/2014 | 87 | Hospital | No |
| West Midlands | 07/06/2014 | 2 | Community | No |
| West Midlands | 07/06/2014 | 62 | Hospital | No |
| West Midlands | 07/06/2014 | 87 | Hospital | No |
| West Midlands | 08/06/2014 | 17 | Hospital | No |
| West Midlands | 08/06/2014 | 43 | Hospital | No |
| West Midlands | 09/06/2014 | 90 | Hospital | No |
| West Midlands | 10/06/2014 | 58 | Hospital | No |
Discussion question 1a
| 1. What further information would you like on the cases and how would you get it?
2. What action would you take? Why? 3. At this point, would you consider this as an outbreak? Why? What measures would you take because it is hospital associated? |
Inject 1b: Whole genome sequencing data on West Midlands PT14b
A week after the isolates making up the West Midlands exceedance are received by SRS, the sequencing data has come back from the Genomic Sequencing Unit at Colindale and has been processed through the Gastrointestinal Bacteria Reference Unit pipeline. One of the outputs of this pipeline is a phylogenetic tree (Figure 2), which places isolates into the imputed phylogenetic context.
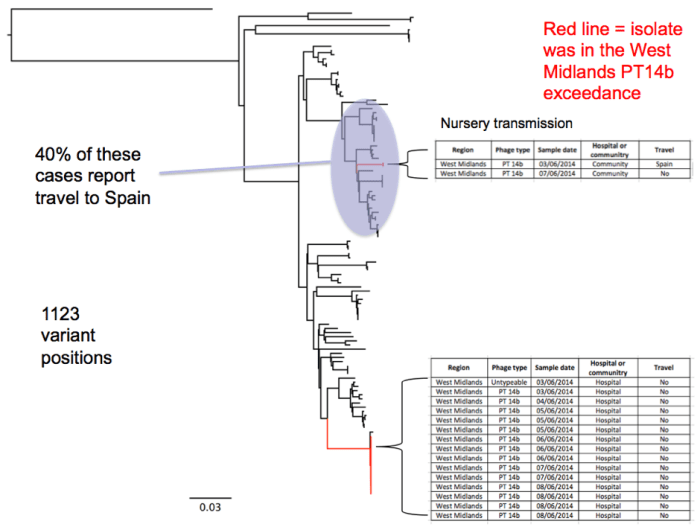
Figure 2: Phylogenetic tree of Salmonella Enteritidis
Discussion question 1b
| Review the information, what conclusions can you draw?
1. Are all the PT14b isolates from the exceedance closely related? 2. What does the WGS tell you about the cases in the hospital? 3. How many SNPs is the outbreak from the most closely related other isolate? What does this suggest about the outbreak? |
Inject 2a: Outbreak goes national
More than a month after the initial West Midlands event and the outbreak has gone national! Over a 3 week period a large number of cases occur in Cheshire & Merseyside and Hampshire & Isle of Wight & Dorset. The clusters in Cheshire & Merseyside and Hampshire & Isle of Wight & Dorset are associated with Chinese restaurants.
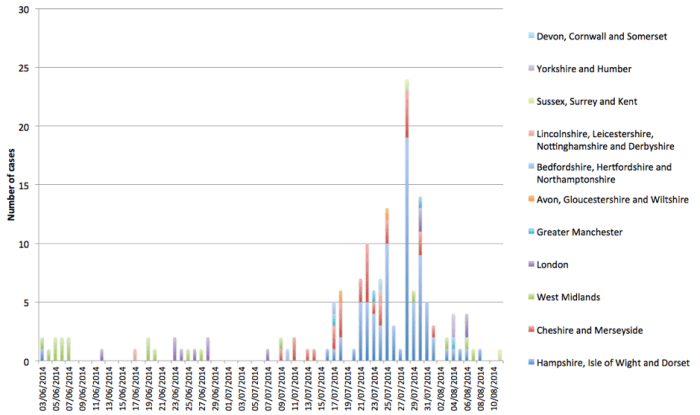
Figure 3: Epidemic curve of outbreak up to present
Table 2: Summary of the number of cases in different regions
| PHE centre | Total number of cases |
| Hampshire, Isle of Wight and Dorset | 82 |
| Cheshire and Merseyside | 29 |
| West Midlands | 18 |
| London | 12 |
| Greater Manchester | 4 |
| Avon, Gloucestershire and Wiltshire | 2 |
| Bedfordshire, Hertfordshire and Northamptonshire | 2 |
| Lincolnshire, Leicestershire, Nottinghamshire and Derbyshire | 2 |
| Sussex, Surrey and Kent | 2 |
| Yorkshire and Humber | 2 |
| Devon, Cornwall and Somerset | 1 |
Discussion question 2a
| 1. How does this change your interpretation of the West Midlands outbreak
2. What do you expect the sequencing results to look like? Consider the pathogen type (i.e. foodborne) and the spatio-temporal scale. |
Inject 2b: Sequencing comes back on the national outbreak
Seven to ten days after the isolates are received by Colindale, we get the sequencing results back and we generate a new phylogeny based on all the isolates (Fig 4).
Figure 4: Phylogenetic tree of national outbreak
Discussion question 2b
| 1. What is your interpretation of the sequencing results? How does it relate to what you predicted the sequencing results to look like?
2. What could explain the differences between the geographically separated outbreaks? 3. Would you say this is a single outbreak? |
Summary exercise
- What difference would having WGS make to your current practice?
- In order to make a difference to your practice, how quickly would you need sequencing info?
- Are there different impacts on practice associated with different levels of timeliness? i.e. 3 days (not currently attainable), 7 days (target), 14 days (maybe just allows a more detailed big picture view).
- Do you feel you are missing events in your region due to insufficient sensitivity of current typing?
- How do you feel about the resource implications of detecting more events compared with not investigating as many false positives?
Potential answers 1a:
- Request molecular typing. Request a review of the historical laboratory data. How many cases were reported in the previous week – is this a continuation of an event?
- Talk to the reference laboratory – are there any other cases coming through the laboratory that may be linked? Do they have any extra information on cases?
- Hard to say at this point in time, it certainly is an increase that requires more investigation. Infection control? Staff screening? Antibiotic resistance?
Potential answers 1b:
- Not all the PT14b isolates are related. There is a main group at the bottom of Fig 2 that is from the hospital, then there are two additional isolates some distance from the main set of PT14bs. This is because phage type is not always congruent with genetic background (i.e. SNPs). The exceedance, which is based on phage type, will group these together, while we can tell from WGS that they do not share a recent common ancestor and should not be treated as part of the same event. The ‘other’ PT14b clade is associated with travel to Spain i.e. 40% of isolates in that clade report travel to Spain. One of these cases was travel to Spain, the other was person to person transmission of the Spain case. Same nursery.
- WGS shows us that all the cases from the hospital are closely related and that there may be a common source of infection. We can also see on the tree that one isolate in the outbreak clade is not PT14b (i.e. red). This isolate was untypeable by phage typing but WGS confirms that it is part of the outbreak.
- The long answer is; all we need to calculate the number of SNPs is two pieces of info. The scale bar on the tree and the number of variant positions. To get the number of SNPs represented by a particular branch, you get the ‘length’ of the branch, in terms of the scale bar. This should be around 0.04, then you multiply that by the number of variant positions i.e. 0.04 * 1123 = 45 SNPs. The short answer is 45 SNPs. This suggests that this outbreak is not closely related to anything else we have ever sequenced.
Potential answers 2a:
- Less emphasis on the role of the hospital? It is community infection that has got into hospital. It raises the importance and the value of traceback analysis as a larger outbreak.
- There is no correct answer to 2. You just need to think about phylogenies in the context of epidemiology. The main ideas people come up with might be a) a big long line, with no variants between the samples b) closely related but separate clusters for each of the geographic locations c) actually unrelated outbreaks, cheshire and mersy/Hampshire are not in the same 14b clade. Summer holidays to Spain may have led to more cases in Spain.
Potential answers 2b:
- This tree reflects a series of point source outbreaks separated by 5-10 SNPs, the original source of which has quite a lot of diversity. How does this relate to what was predicted in 2a?
- There are more SNPs between these point source outbreaks than you would expect if they all shared a recent common ancestor. What could explain this? The accumulation of a large number of mutations in a short space of time is not likely. More likely to be sampling from a diverse source.
- Genetically and temporally clustered which indicates yes. Geographical distribution implies no? Need to put sequencing into the context of time-place-person epidemiology. With any other typing, would be monomorphic.
Appendix 1
Glossary of terms
- WGS: whole genome sequencing, determining the sequence of DNA encoded by an organism.
- SNP: Single nucleotide polymorphism, the most common type of variant used in phylogenetic analyses arising from whole genome sequencing
- Phylogenetics: the study of evolutionary relationships among groups of organisms through study of biological macromolecules (e.g. DNA).
- Case-case analysis: An epidemiological method to compare cases in an outbreak to cases of similar disease who are not part of an outbreak: http://ije.oxfordjournals.org/content/28/4/764.full.pdf?origin=publication_detail
- ECDC: European Centre for Disease Prevention and Control: http://www.ecdc.europa.eu/en/Pages/home.aspx
- EHO: Environmental Health Officer: http://www.cieh.org/professional_development/becoming_an_EHP.html
- Exceedance: An automated report based on laboratory reporting of human isolates which lists organisms for which more isolates have been reported in the previous week than would be expected based on historical data.
- FSA: Food Standards Agency: http://www.food.gov.uk/
- FWE: Food, Water and Environmental Microbiology Services:
- GEZI: Gastrointestinal, Emerging and Zoonotic Infections department, PHE
- GBRU: Gastrointestinal Bacteria Reference Unit
- OCT: Outbreak Control Team: Set up to coordinate outbreak investigations.
- PT: Phage type: A method of discriminating based on how isolates react to bacteriophages.
- Trawl: A very detailed questionnaire used to inform hypothesis generation.
- SRS: Salmonella Reference Service, the national reference lab for Salmonella
- Clade: a monophyletic group, may be very closely related or more distantly related.
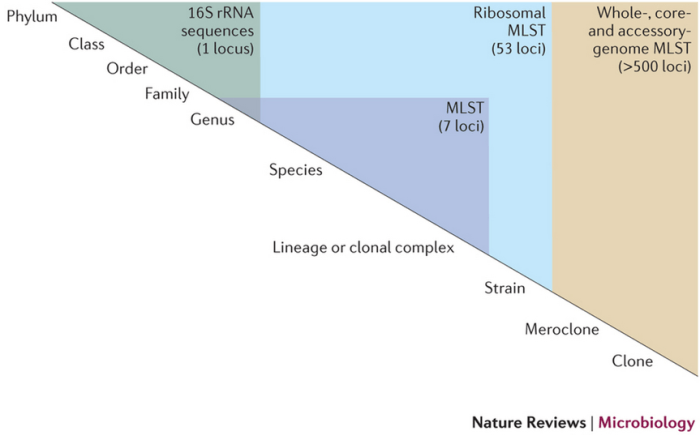
Appendix 2
Supp figure 1: A nomenclature for discussing the genetic relatedness of bacteria. Maiden et al., Nature Reviews Microbiology 11, 728–736 (2013) doi:10.1038/nrmicro3093
Other sources of information:
- Youtube video of a presentation on the Salmonella WGS work at PHE: http://bit.ly/1M8PZWm
- List of WGS papers that are relevant to public health microbiologists and epidemiologists: http://bit.ly/1ALLk6l